CellMLAnnotationView plugin¶
The CellMLAnnotationView plugin can be used to annotate CellML files by associating one or several ontological terms to a given CellML element. To open a CellML file that does not contain any annotations gives:
All the CellML elements that can be annotated are listed on the left of the view. Right click on any of them to get a popup menu, which you can use to expand/collapse all the child nodes, as well as remove the metadata associated with the current CellML element or any CellML element in the CellML file:
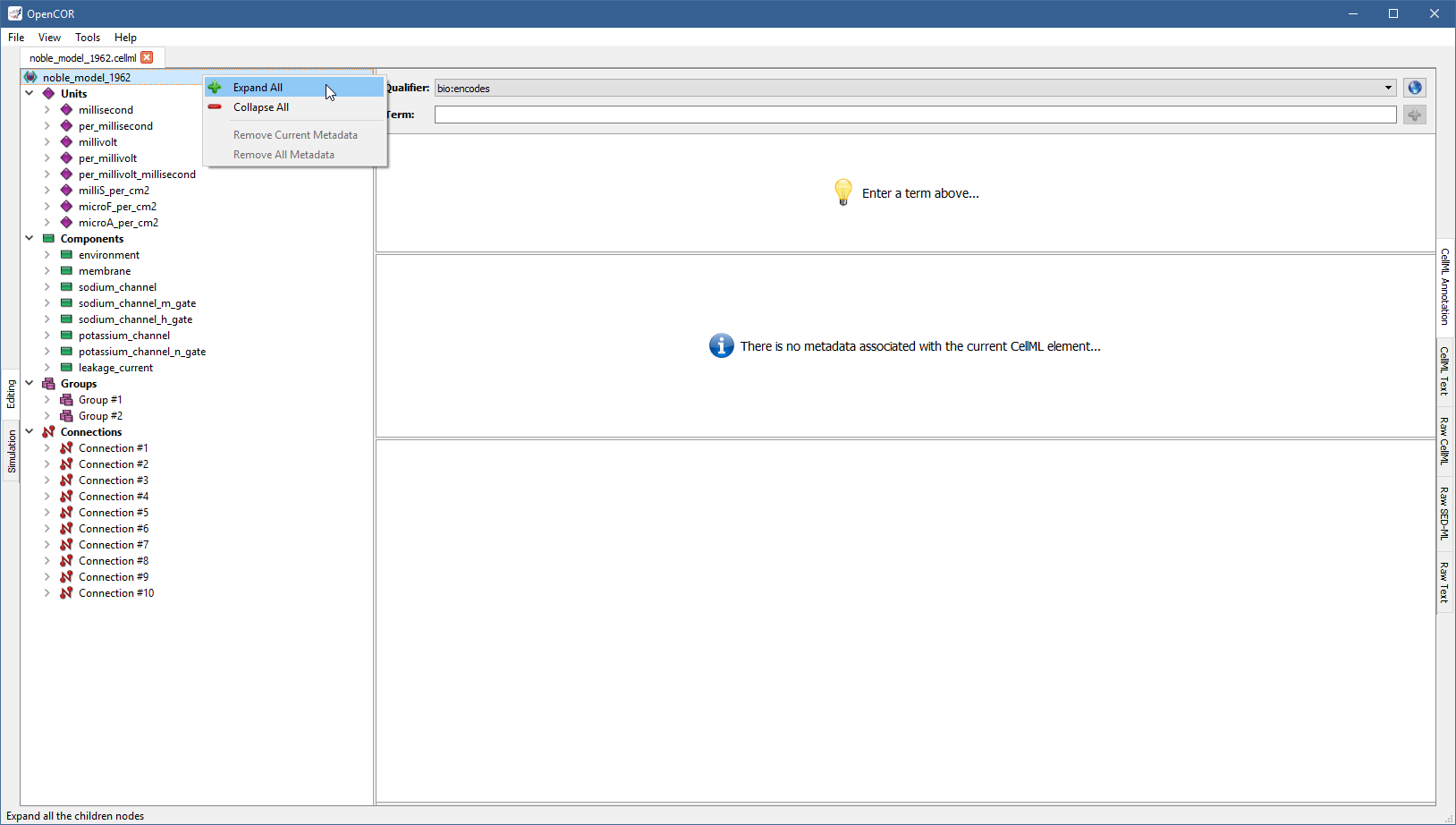
Annotate a CellML element¶
To annotate, say, the sodium_channel component, you first need to select it:
Next, you need to specify a BioModels.net qualifier.
If you do not know which one to use, click on the  button to get some information about the current BioModels.net qualifier:
button to get some information about the current BioModels.net qualifier:
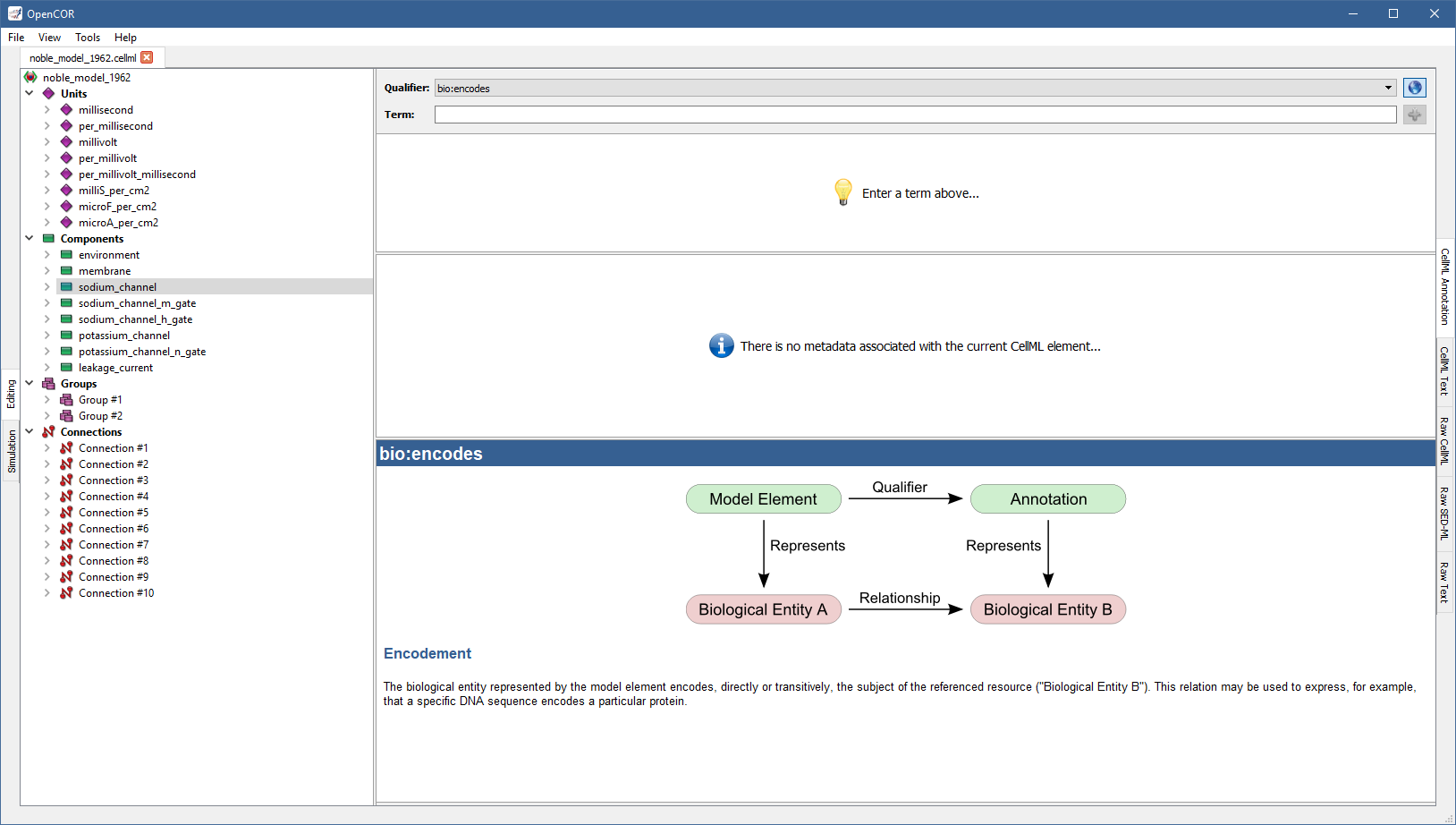
From there, go through the list of BioModels.net qualifiers until you find the one that suits your needs best (e.g., bio:isVersionOf):
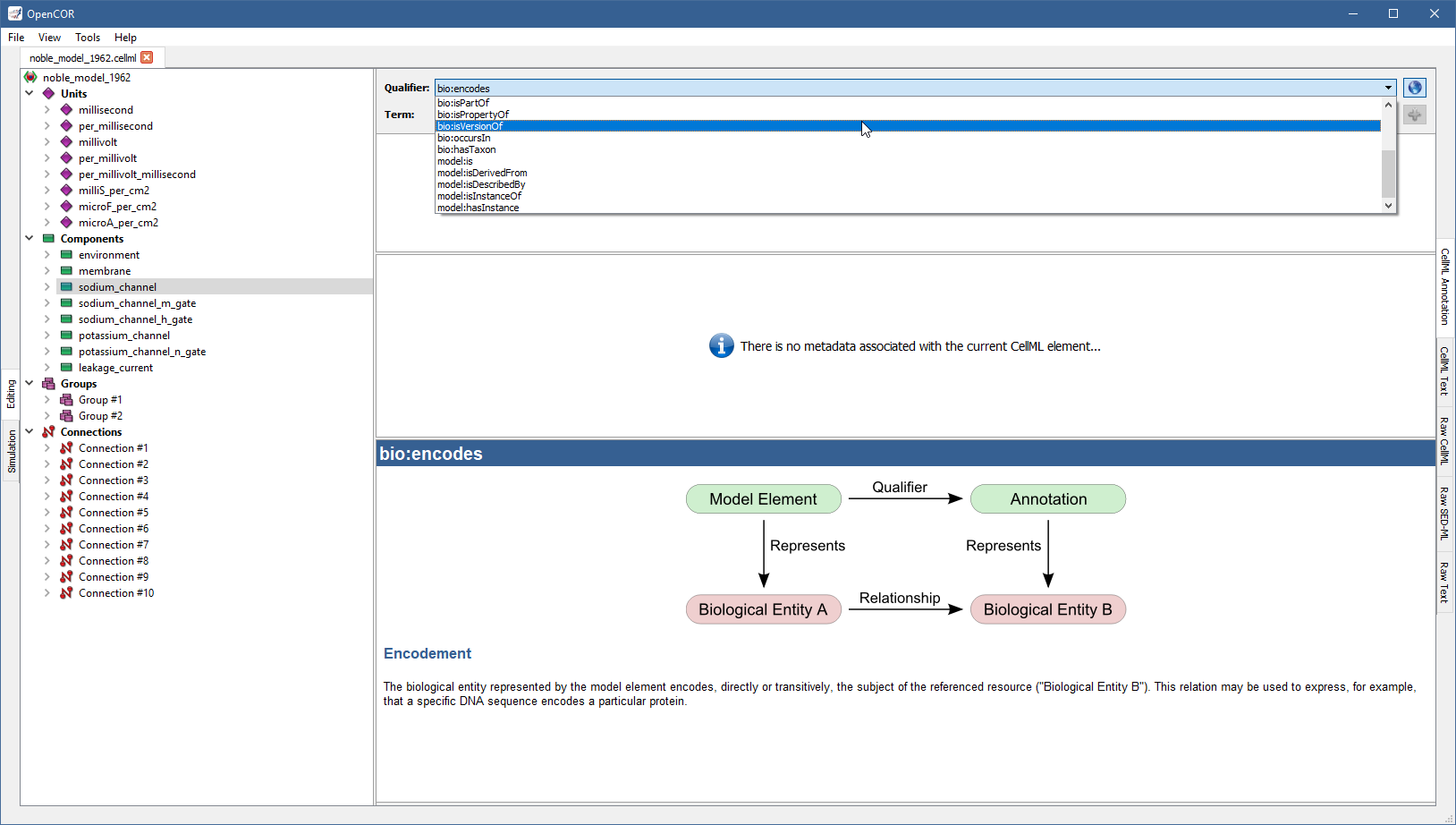
Now, you need to retrieve some possible ontological terms to describe the sodium_channel component.
For this, you need to enter a search term (e.g., sodium channel; note: regular expressions are supported).
As can be seen, OpenCOR returns 12 possible ontological terms:
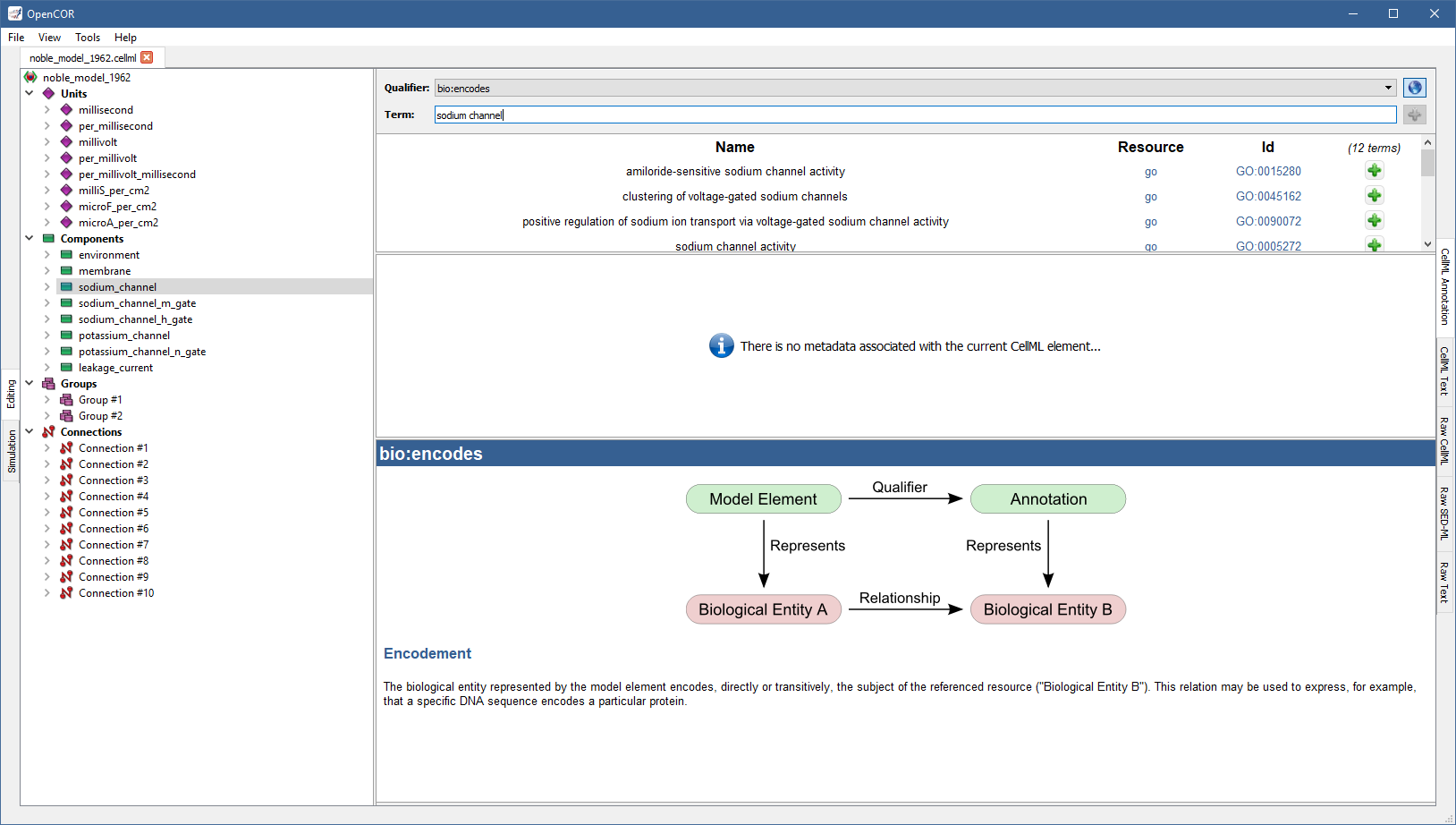
A quick look through the list tells us that you might want to use the one for voltage-gated sodium channel complex.
If you want to know more about the GO resource, you can click on the corresponding link:
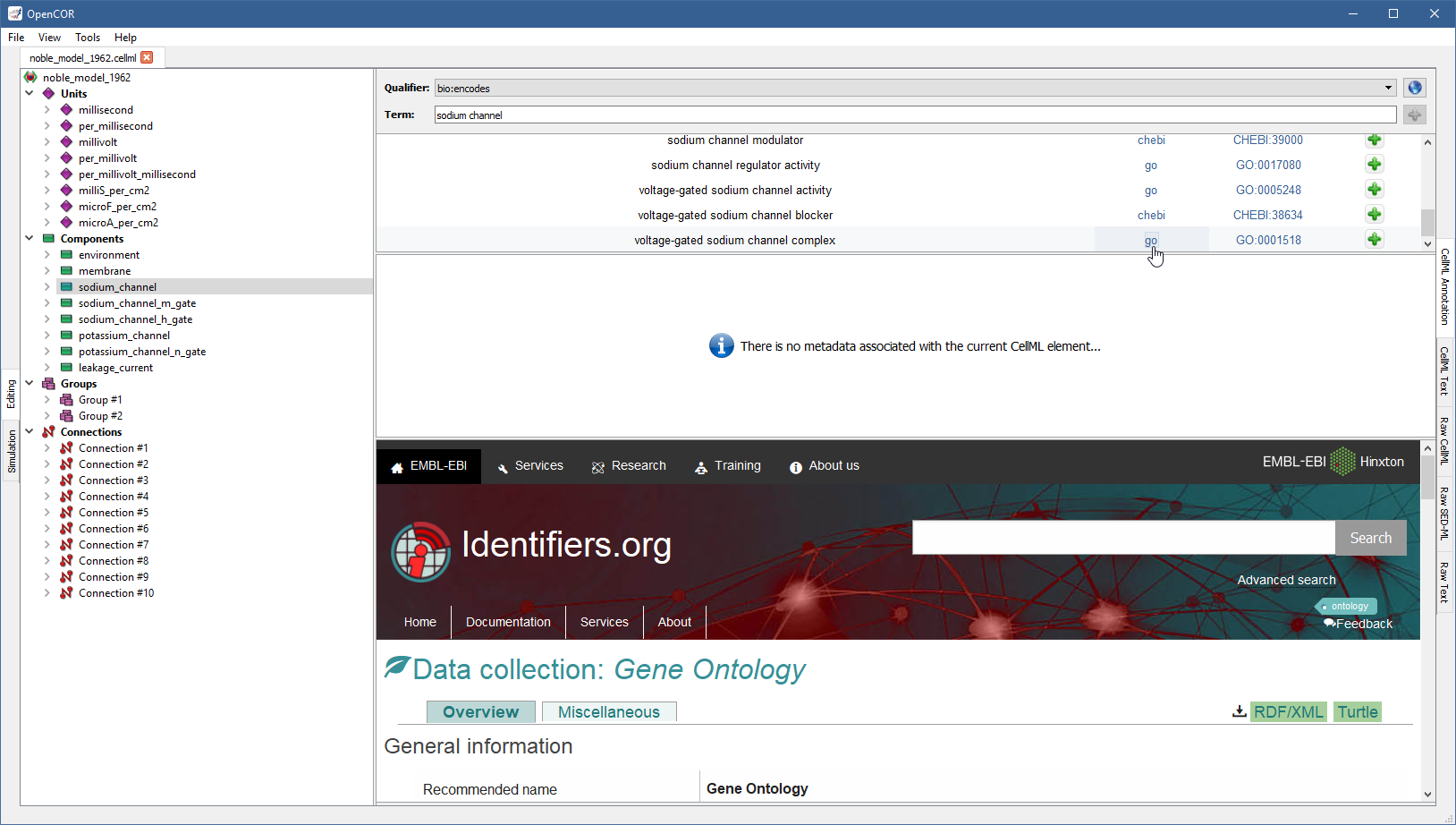
Similarly, if you want to know more about the GO identifier:
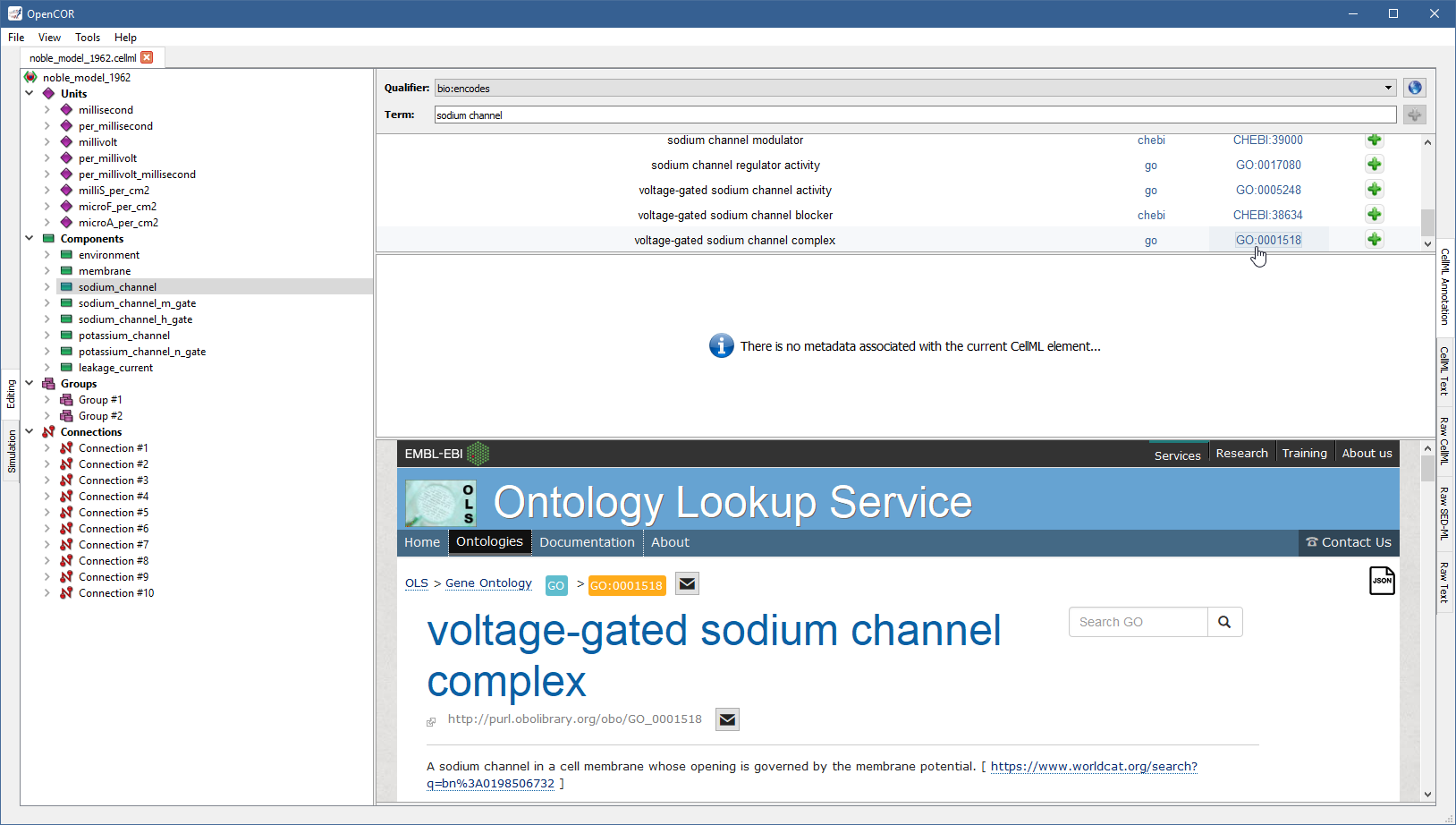
Now, assuming that you are happy with your choice, you can associate the ontological term with the sodium_channel component by clicking on the corresponding  button:
button:
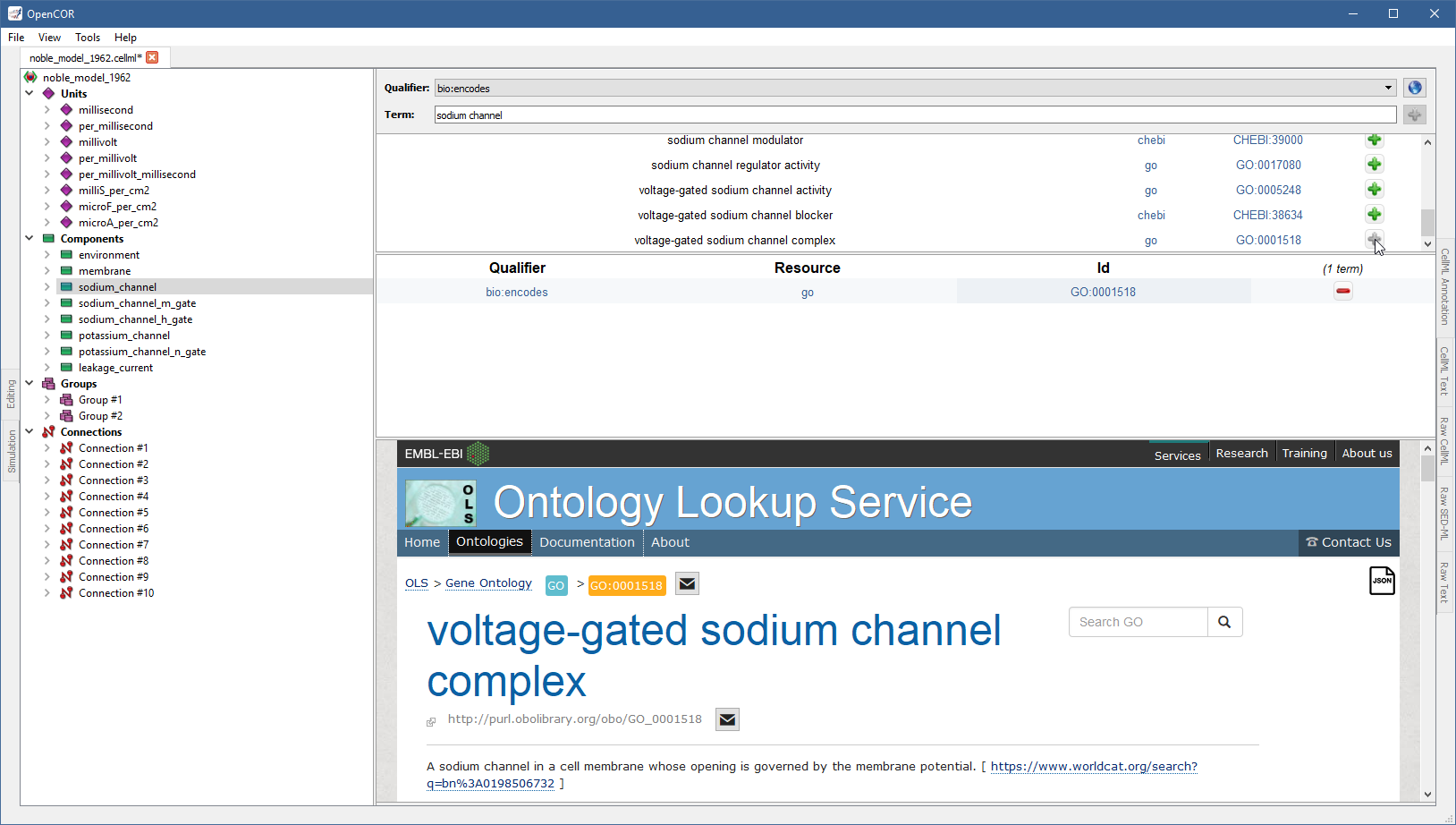
The ontological term you have added cannot be added anymore, but it can be removed by clicking on the corresponding  button.
button.
Now, say that you also want to add another ontological term.
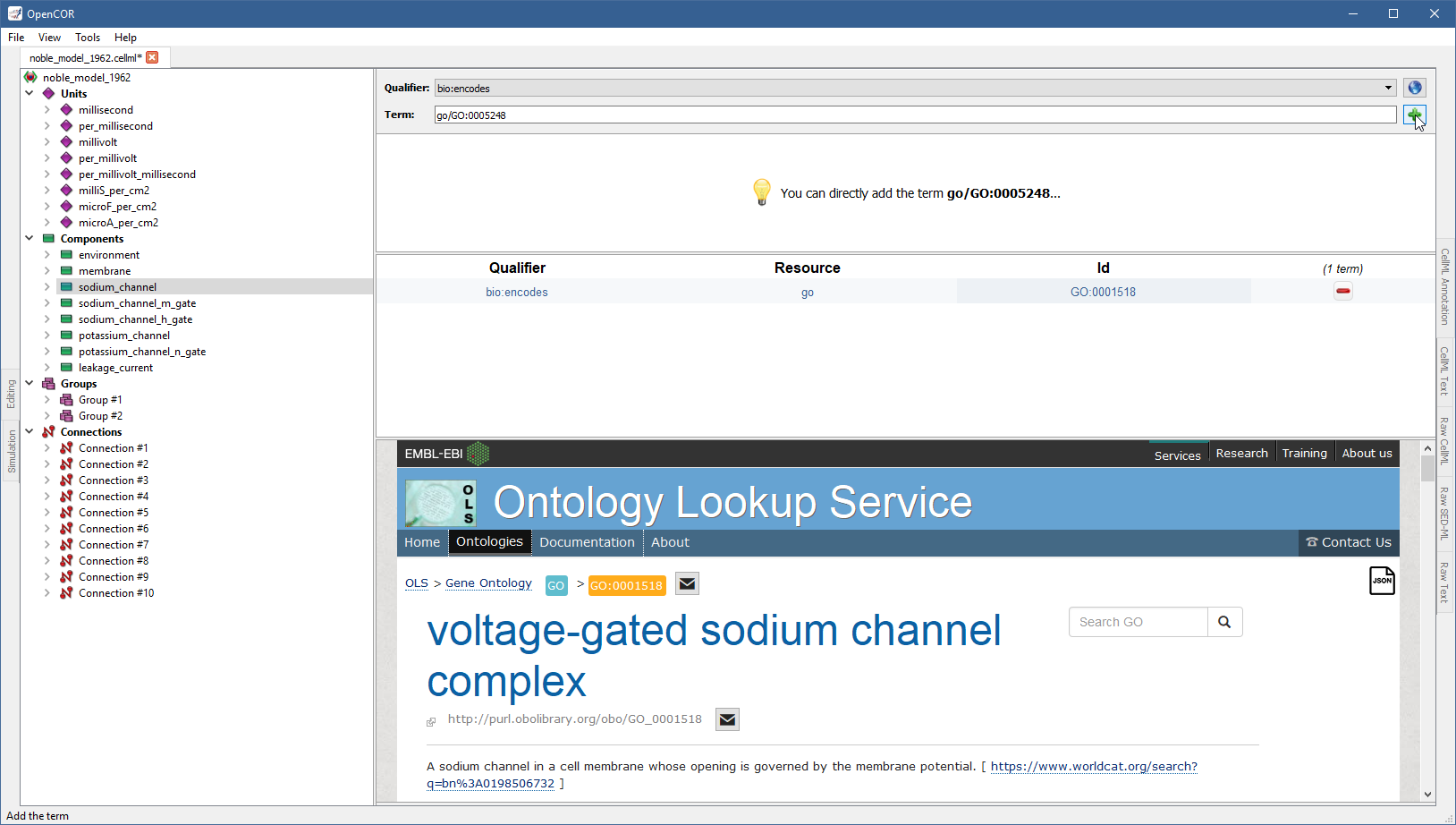
You can do so by clicking on the corresponding  button, but also by entering its resource-id duple (e.g.,
button, but also by entering its resource-id duple (e.g., go/GO:0005248; i.e. <resource>/<id>) in the term field.
OpenCOR will recognise this “term” as being a resource-id duple and will offer you to add the corresponding ontological term directly:
Unrecognised annotations¶
Annotations consist of RDF triples, which are made of a subject, a predicate and an object. OpenCOR recognises RDF triples, which subject identifies a CellML element while it expects the predicate to be a BioModels.net qualifier and the object an ontological term.
Ontological terms used to be identified using MIRIAM URNs, but these have now been deprecated in favour of identifiers.org URIs. OpenCOR recognises both, but it will only serialise annotations using identifiers.org URIs.
Now, it may happen that a file contains annotations that are not recognised by OpenCOR. In this case, OpenCOR will display the annotations as a simple list of RDF triples:

If you ever come across such a type of annotations and think that OpenCOR ought to recognise it then please do get in touch.